Dinoflagellates constitute an important role in the primary productivity of the world oceans, with numerous species have been identified responsible for harmful algal blooms (HABs) and poisoning events in coastal areas. Although, studies of marine dinoflagellates, their distributions and harmful effects are increasing worldwide, little is known about the biodiversity of dinoflagellate assemblages in the Saudi Arabian Red Sea, especially the benthic and toxic forms. Previous studies have highlighted the importance of dinoflagellate cysts as bio-indicators for environmental conditions. Among other factors, the formation of resting cysts has often been associated with survival of dinoflagellates during unfavourable environmental conditions: e.g. changes in temperature and salinity, light radiation, nutrient limitation, low oxygen, etc. Hence, a baseline is required if one is to follow responses in biodiversity, distributions, seasonal dynamics and harmful effects of Red Sea dinoflagellates to global change like ocean warming, UV radiation, nutrients limitation, pollutants and other stressors. The hypothesis underlying this study is that the oligotrophic conditions of the Red Sea are suspected to have influenced the evolutionary adaptation of dinoflagellates in this region, making them not only distinct but able to tolerate changes in environmental better than other species found from different regions.
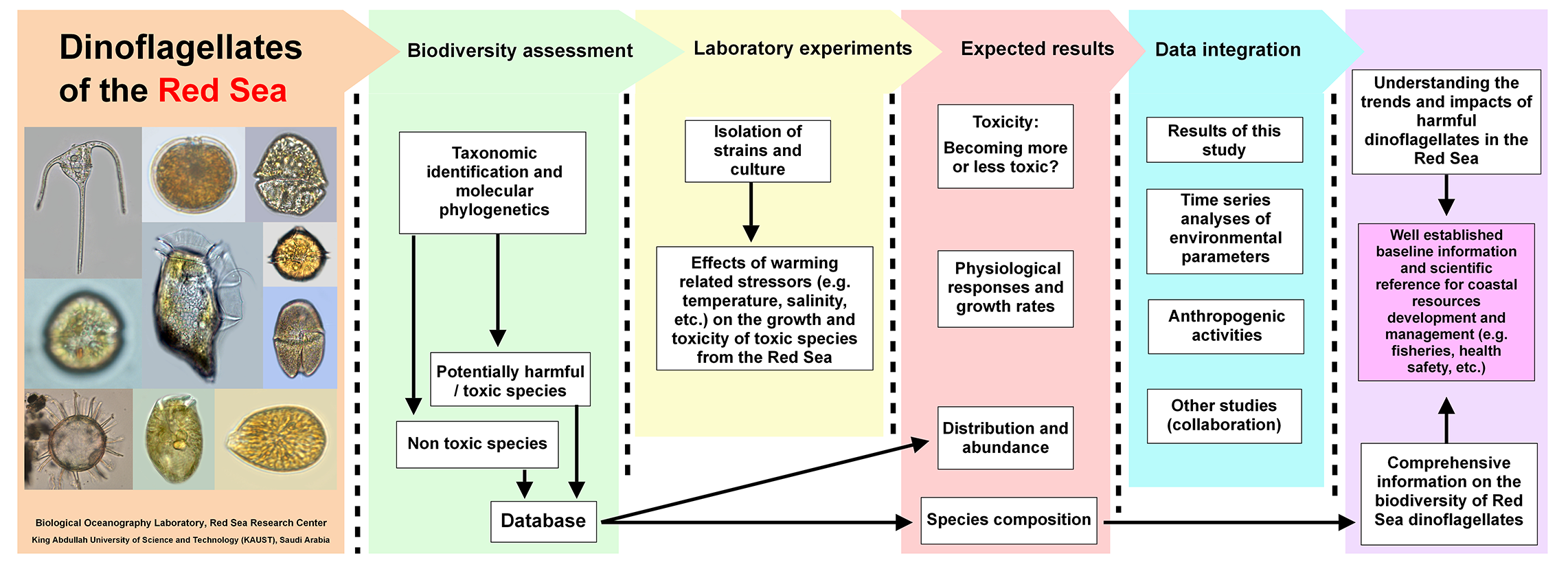
Our study aims to unveil hidden biodiversity of Red Sea dinoflagellates with special emphasis on harmful/toxic species and investigate their responses to warming related stressors. Furthermore, the potentiality of discovering novel dinoflagellate species from this area is considered. The knowledge acquired in this study, combined with the information obtained from the time series analyses of environmental parameters in this area, is expected to rise the interest of local authorities towards the protection and monitoring of Red Sea`s coastal resources to anticipated harmful dinoflagellates blooms, poisoning events and environmental changes in the future. To meet this aim, the following specific objectives are conducted:
- To isolate and culture potentially harmful dinoflagellate strains from the Red Sea
- To perform taxonomic / systematic identification (i.e. based on morphology and molecular markers) and toxicity analyses of dinoflagellate species and use these data to contribute to the known diversity of the area
- To test the responses of harmful / toxic dinoflagellates of interest to stressors (e.g. temperature)
- To make this data available for continuous monitoring of harmful dinoflagellates community in this region
Arwa Aynousah (MSc student) is currently investigating the effects of increased temperature in stimulating the blooms and toxicity of
Prorocentrum sp. from the central Red Sea
There are increased attention in studying the correlation between climate change and harmful algal blooms (HABs). Some studies had discovered that climate change might be one of the factors involved in stimulating the growth and toxicity of dinoflagellates. Such factors could be physical parameters (i.e. temperature, salinity, and light) or the availability of certain nutrients in seawater. However, to the best of our knowledge, there are limited studies conducted about the effects of climate change and HABs (i.e. toxic dinoflagellates), especially in the Red Sea. For our study, we are interested on Prorocentrum spp. isolated from the central Red Sea. Some marine benthic and epiphytic Prorocentrum spp. are responsible for diarrheic shellfish poisoning (DSP) because they produces okadaic acid (OA) and dinophysis toxins (DTXs). We hypothesized that there is positive correlation between the increase of temperature and the growth as well as toxin production of Prorocentrum species in the Red Sea. To be able to proove our hypothesis, we will employ multiphasic investigation involving experiments on microalgal cultures, microscopic observation, molecular analyses and analytical chemistry (i.e. HPLC-MS) using state-of-the-art technologies supported by KAUST Core Lab.